African swine fever virus interactome (ASFVint)
ASFVint is a major international collaborative project to understand the breadth, depth and impact of African swine fever virus' interactions with its host.
A major challenge in developing strategies to fight African swine fever virus is the lack of knowledge of how viral proteins interact with the proteins of the cells which it infects. Cells are complex structures that contain thousands of different proteins that are involved in a myriad of tasks ranging from the spatial organization and shape of the cells, production of new proteins, sugars, lipids, nucleic acids as well as any housekeeping and maintenance. Proteins also play a vital role in communicating within and between different cells and cell types.
Viruses are obligate parasites that need to harness cells and their proteins in order to successfully make copies of themselves and survive. Whenever a virus infects cells a struggle arises between the cellular or host proteins and their viral counterparts. With thousands of host proteins and multiple viral proteins these networks of interactions can become incredibly complex and the science of interactomics is how biologists try to solve this problem.
Our research
Understanding how viral proteins target cellular functions will drive development of vaccines and antivirals.
The African swine fever virus interactome (ASFVint) project will use high-throughput systems biology approaches to map interactions between viral and cellular proteins in a systematic way. Six partners from leading research institutes in Europe will combine their expertise in African swine fever virus and protein-protein interactions to define and validate the first African swine fever virus interactome.
The map (below) shows the known protein-protein interactions for African swine fever virus at the beginning of the project in 2021 compared to a similarly sized virus called herpes simplex virus.
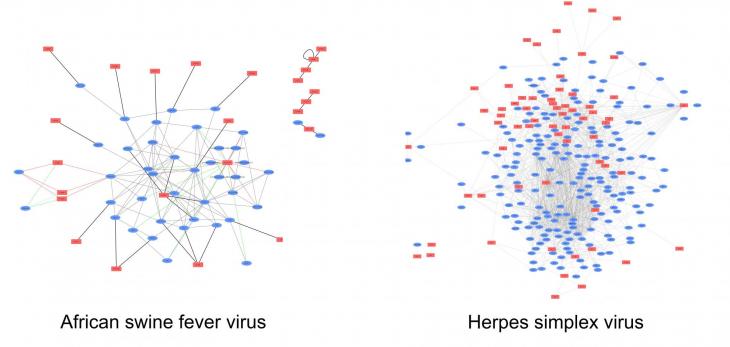
By the end of our project we aim to make an African swine fever virus interactome map to the same or a better standard than the herpes simplex virus map. We'll do this using three complementary approaches.
Breadth
African swine fever virus has more than one hundred and fifty genes and we only understand the role of a few of these in the life cycle of the virus. Looking at all of these genes at once would take be a massive undertaking so we will look for pig proteins that interact with more than eighty viral proteins that we have selected using several overlapping approaches. This will provide the basic outline of the map that we will use to build the African swine fever virus interactome.
Depth
We will take a deep dive into some of the interactions of pig and viral proteins. Can we detect direct interactions between pairs of proteins in pig cells? Which parts of the proteins interact with each other? Can we detect protein-protein interactions involving multiple proteins?
Impact
We will test the importance of interactions during African swine fever virus infection by looking at the level of the host cell and the virus itself. What effect does modifying or deleting host genes effect the ability of the virus to replicate in the cell? How does modifying or deleting viral genes have on the virus? Do these changes affect the ability of the virus to disrupt the cell's pathways? Are any cellular proteins essential for the virus to make copies of itself?
Who we are
| Ploufragan-Plouzané-Niort Laboratory at Agence Nationale de Sécurité Sanitaire de l'Alimentation, de l'Environnement et du Travail (ANSES) in France. |  |
| African swine fever vaccinology and African swine fever virus research groups at The Pirbright Institute in the United Kingdom. |  |
| Laboratory for Biochemistry and Protein Analysis at Friedrich-Loeffler-Institut (FLI) in Germany. |  |
| The Biology of Orbiviruses Team at Institut National de Recherche pour l’Agriculture, l’Alimentation et l’Environnement (INRAE) in France | |
| The Biotechnology Department of Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria (INIA) in Spain. |  |
| Bioinformatics, Algorithmics and Data Mining Group at the Institute of Computer Science of the University of Tartu in Estonia. |
We are funded by our national agencies through the International Coordination of Research on Infectious Animal Diseases (ICRAD).
Project Abstract
A major challenge in the development of strategies to prevent and fight ASFV is the absence of knowledge of the functions of viral proteins and their interactions with host proteins which is a prerequisite for the rational development of new antiviral strategies, or vaccines. Functional characterization of virus-host protein-protein interactions will be critical to understand how viral proteins target cellular functions to allow viral replication and spread. To achieve this goal, we will use high-throughput systems biology approaches to map interactions between viral and cellular proteins in a systematic way. In this project, six partners from leading research institutes (ANSES, FLI, INIA, INRAE, The Pirbright Institute and UTARTU) will combine their expertise in ASFV and protein-protein interactions to define and validate the first ASFV interactome.
As a starting point, a high-throughput yeast two-hybrid (Y2H) approach will be performed using 80 selected viral proteins encoded by ASFV as baits to screen a cDNA library from domestic pig macrophages, the target cells for ASFV replication. An affinity purification mass spectrometry (AP-MS) method will also be used to extend our functional interactomic strategy in a multiparametric analysis. Data obtained from our laboratories and others have provided evidence that ASFV targets the host interferon (IFN) and autophagy responses which could play a significant role in ASFV virulence and pathogenesis. Therefore, in parallel to these unbiased proteomic approaches, the function of selected ASFV proteins in IFN and autophagy cellular pathways will be studied using multiple approaches. The most relevant interactions will be validated by co-immunoprecipation, AP-MS, and NanoLuc assays and characterized in detail by microscale thermophoresis. After confirmation of the physical interactions by biochemical approaches, their biological significance will be independently tested in the context of infection by deleting genes of interacting viral proteins and/or silencing interactors from the host. Using this strategy we will evaluate the potential pro or antiviral functions associated with the interactions identified. We have the unique advantage of the tools and skills from partners within this consortium in the genetic manipulation of ASFV and the benefit of already having a number of gene-deleted ASFV strains available in our laboratories. Such recombinant viruses and strains, in comparison with their corresponding parental/wild type viruses, will be very valuable to study the role of a specific protein interaction in viral replication.
The ASFVint project is designed to identify cellular signalling pathways, functional modules, and machineries that are manipulated by the virus to its own benefit or even are essential for ASFV replication. Knowledge of such pathways would represent a valuable resource for the development of antiviral strategies. Collectively, deciphering virus-host molecular interactions opens new perspectives to predict/simulate future emergences and develop effective countermeasures for disease control, such as novel anti-infectious compounds and rationally designed ASFV vaccines.

