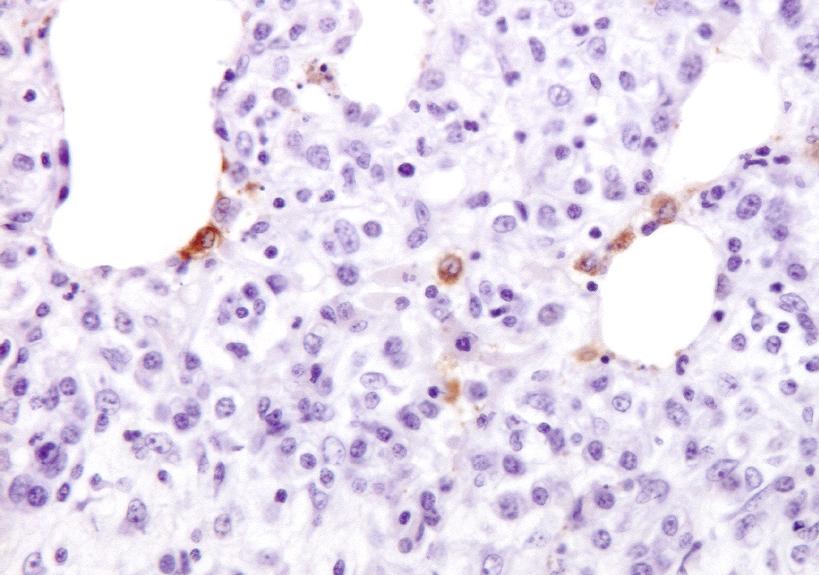
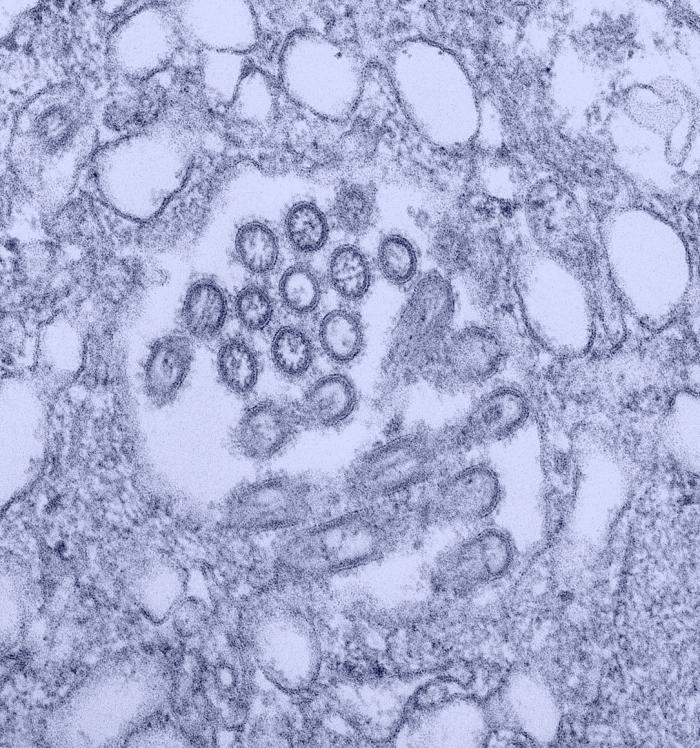
Changes in the nasal microbiota of pigs following single or co-infection with porcine reproductive and respiratory syndrome and swine influenza A viruses
Host-microbiota interactions are important in shaping immune responses that have the potential to influence the outcome of pathogen infection. However, most studies have focused on the gut microbiota and its possible association with disease outcome, while the role of the nasal microbiota and respiratory pathogen infection has been less well studied. Here we examined changes in the composition of the nasal microbiota of pigs following experimental infection with porcine reproductive and respiratory syndrome virus 2 (PRRSV-2), swine influenza A H3N2 virus (H3N2) or both viruses. DNA extracted from nasal swabs were subjected to 16S rRNA sequencing to study the composition of the nasal microbiota. Bacterial richness fluctuated in all groups, with a slight reduction in pigs singly infected with PRRSV-2 and H3N2 during the first 5 days of infection compared to uninfected controls. In contrast, nasal bacterial richness remained relatively stable after PRRSV-2/H3N2 co-infection. PRRSV-2 and H3N2, alone or in combination differentially altered the abundance and distribution of bacterial families. Single and co-infection with PRRSV-2 or H3N2 was associated with the expansion of the Neisseriaceae family. A positive correlation between H3N2 viral load and the relative abundance of the Neisseriaceae was observed. However, further mechanistic studies are required to understand the significance of the changes in specific bacterial families following these viral infections.