The African swine fever virus transcriptome
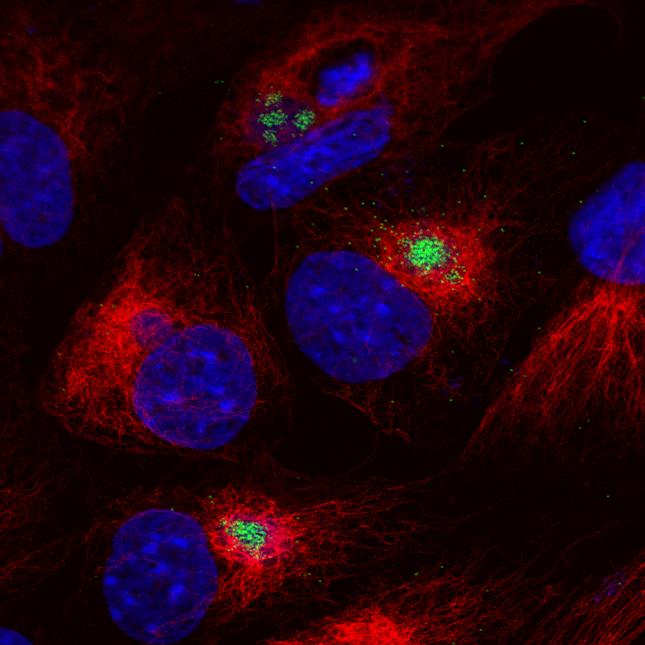
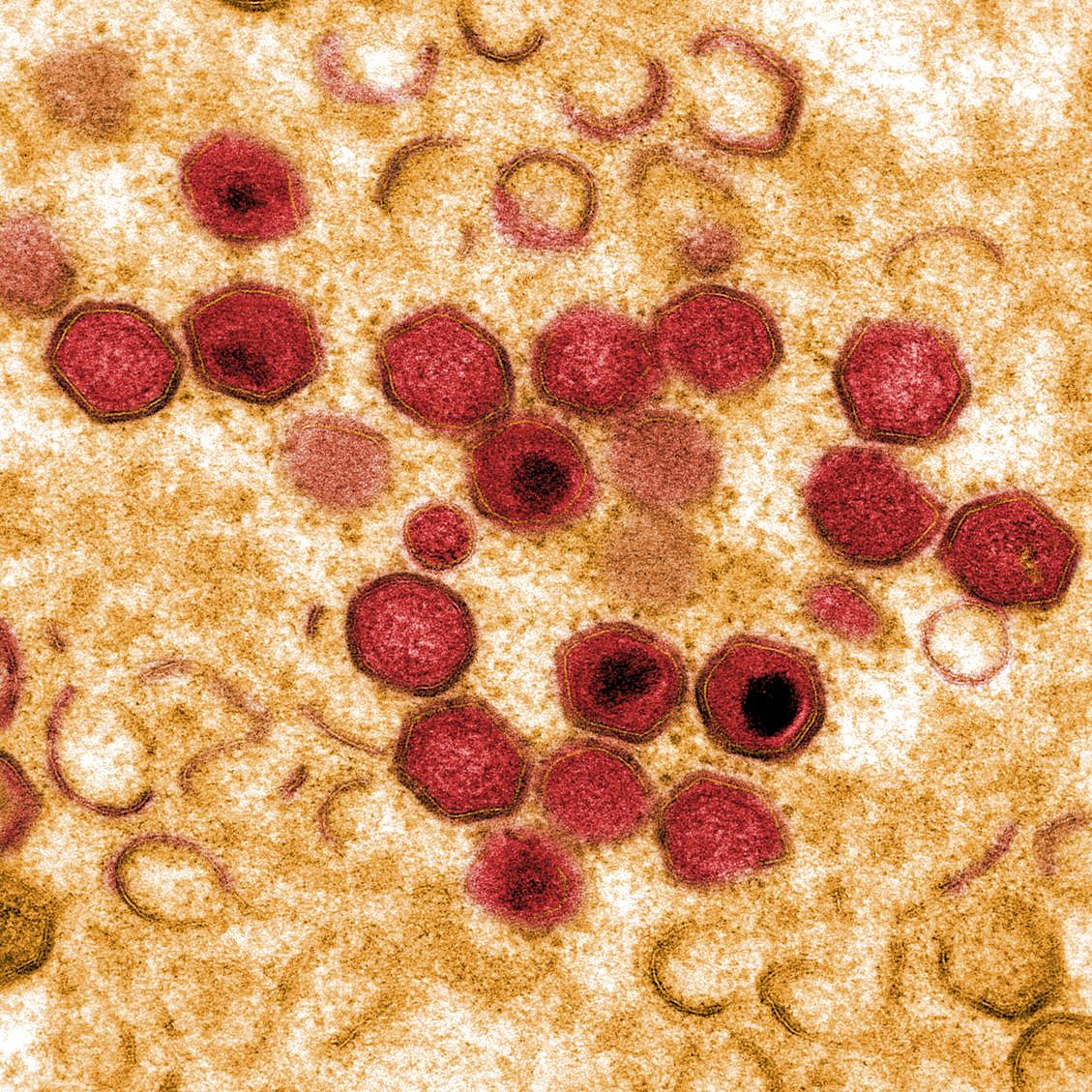
African Swine Fever Virus (ASFV) causes haemorrhagic fever in domestic pigs, presenting the biggest global threat to animal farming in recorded history. Despite its importance, little is known about the mechanisms and regulation of ASFV transcription. Using RNA sequencing methods, we have determined total RNA abundance, transcription start sites and transcription termination sites at single nucleotide-resolution. This allowed us to characterise DNA consensus motifs of early and late ASFV core promoters, as well as a poly-thymidylate sequence determinant for transcription termination. Our results demonstrate that ASFV utilises alternative transcription start sites between early and late stages of infection, and that ASFV-RNAP undergoes promoter-proximal transcript slippage at 5′ ends of transcription units, adding quasi templated AU- and AUAU-5′ extensions to mRNAs. Here we present the first much-needed genome-wide transcriptome study that provides unique insight into ASFV transcription and serves as a resource to aid future functional analyses of ASFV genes which are essential to combat this devastating disease.ImportanceAfrican swine fever virus (ASFV) causes incurable and often lethal haemorrhagic fever in domestic pigs. In 2019, ASF presents an acute and global animal health emergency that has the potential to devastate entire national economies as effective vaccines or antiviral drugs are not currently available (Food and Agriculture Organization of the UN). With major outbreaks ongoing in Eastern Europe and Asia urgent action is needed to advance our knowledge about the fundamental biology of ASFV, including the mechanisms and temporal control of gene expression. A thorough understanding of RNAP and transcription factor function, and the sequence context of their promoter motifs, as well as accurate knowledge of which genes are expressed when and the amino acid sequence of the encoded proteins, is direly needed for the development of antiviral drugs and vaccines.