Our group
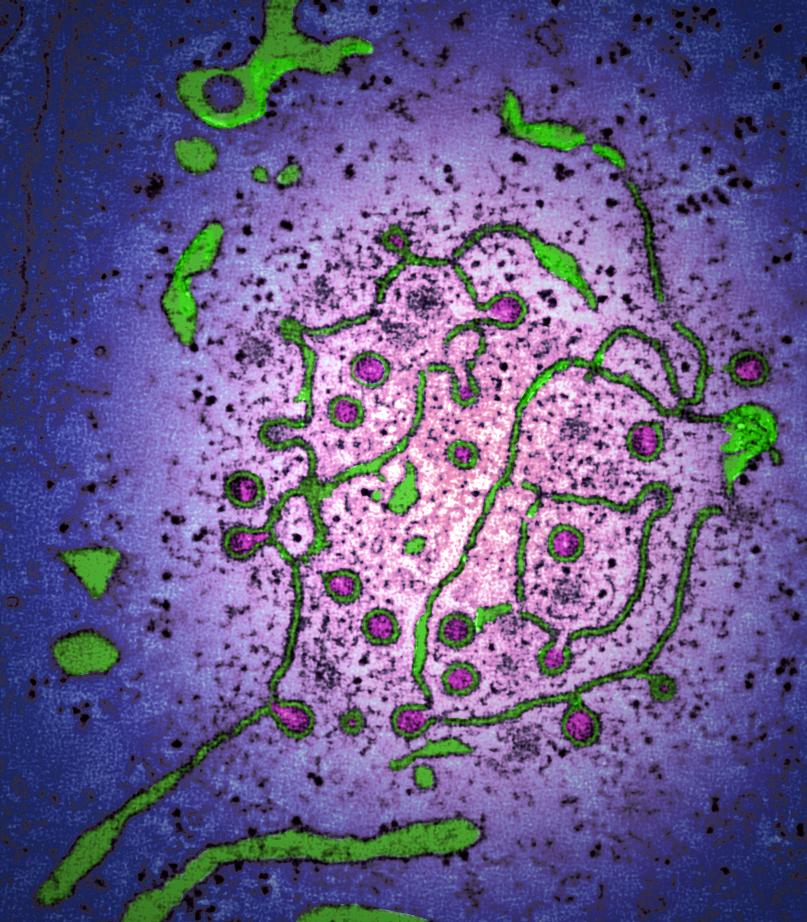
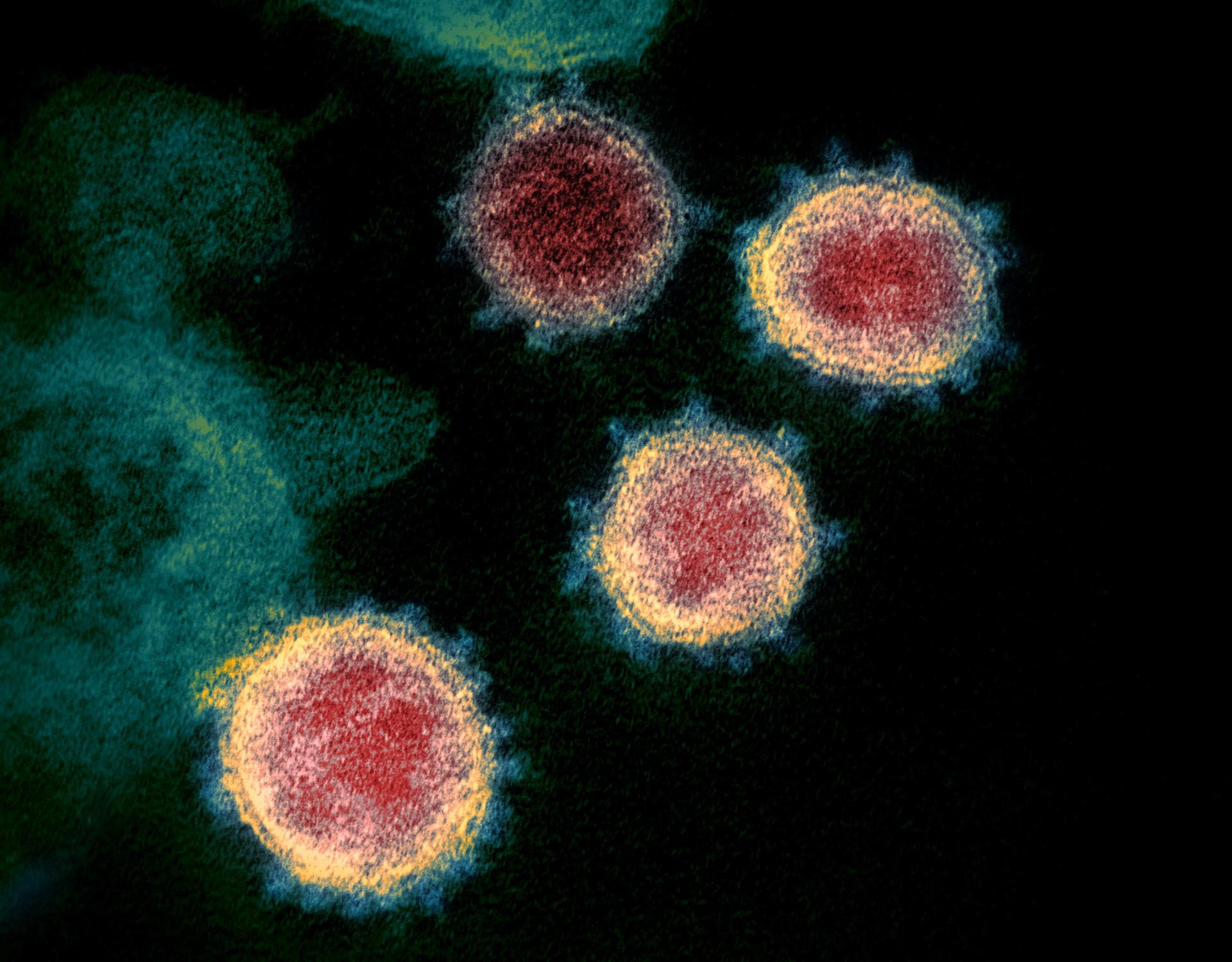
The coronavirus family contains a range of important viruses. These viruses can jump between host species and as shown with the COVID-19 pandemic, can have a significant impact on human health and our way of life. In addition, coronaviruses impact animal species, in particular poultry and pigs, causing disease, affecting food security, and having economic impacts on livestock industries.
The Coronavirus Cellular Biology group, led by Dr Helena Maier, studies molecular interactions between coronaviruses and their host cells, using a comparative approach to identify interactions conserved across the family.
Our aims
The overall aim of the group is to identify critical virus-host cell interactions that are conserved among coronaviruses, ultimately allowing for development of novel approaches for vaccine attenuation or development of pan-antiviral therapeutics. Using advanced bioimaging, multi ‘omics, reverse genetics and cellular and molecular biology techniques our work focusses on understanding replication and cellular interaction of a range of coronaviruses. These include infectious bronchitis virus (IBV), porcine deltacoronavirus (PDCoV), mouse hepatitis virus (MHV), SARS-CoV-2 and human common cold coronaviruses 229E and OC43.
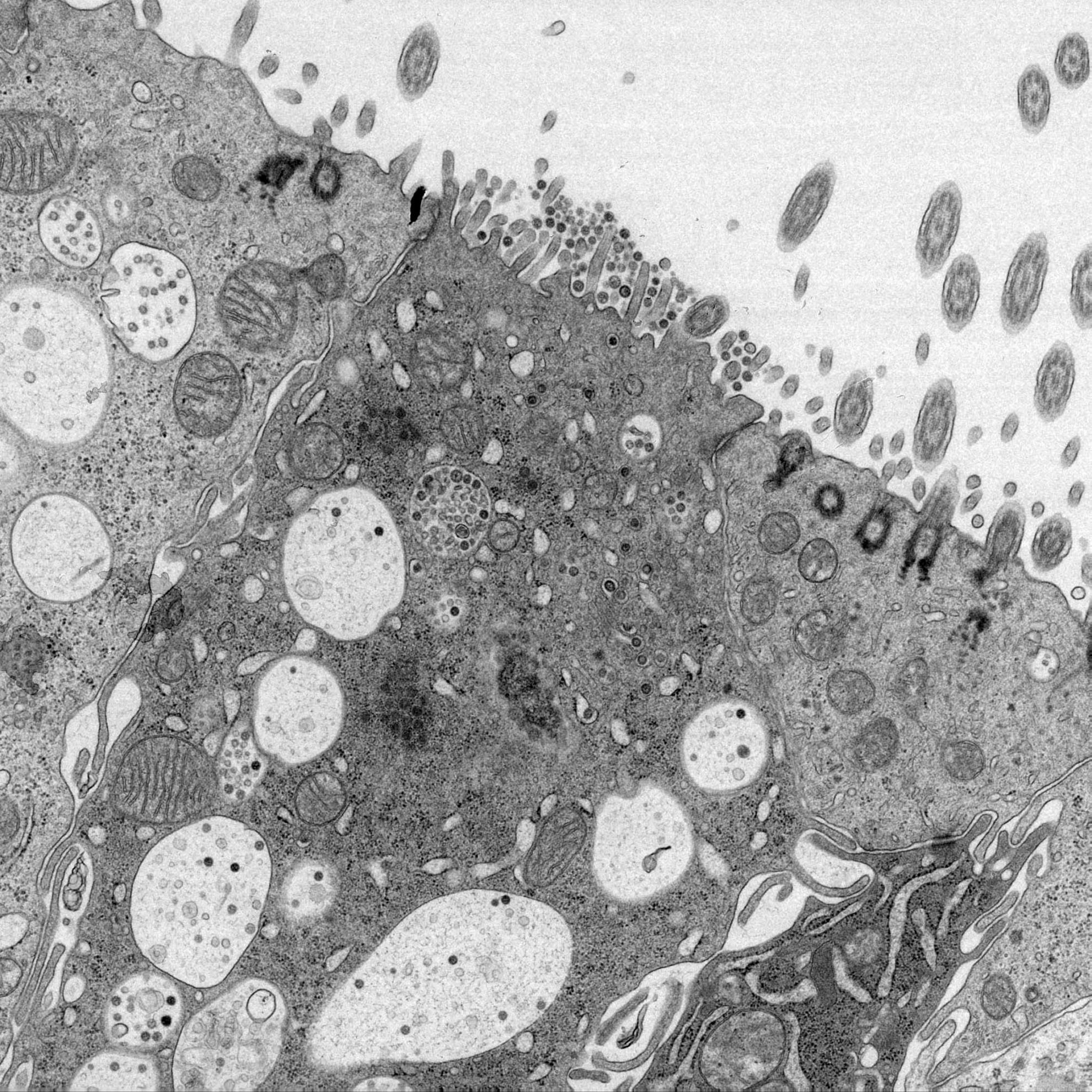
The group has a particular focus on characterising virus induced replication organelles, understanding viral regulation of cellular translation and stress pathways, the process of viral envelopment and identifying cellular proteins that are important for virus replication.
Our research
Our research includes:
- Identification of cellular proteins involved in replication organelle formation and function and conservation across coronaviruses
- Characterisation the structure of coronavirus replication organelles using cryo-soft x-ray tomography and cryo-electron tomography
- Identification of viral proteins responsible for the formation of different elements within the replication organelle
- Identification of essential host factors for the replication of infectious bronchitis virus and infectious bursal disease virus
- Characterisation of coronavirus regulation of antiviral and stress signalling pathways and the role of signalling regulation in cross species transmission
- Characterisation of the role of cellular RNA binding proteins in coronavirus replication
- Identification of cellular proteins required for assembly, budding and exit of coronavirus virions
- Establishment of reverse genetics systems to facilitate study of coronavirus-host cell interactions
Our impact
Coronaviruses are important disease-causing viruses, with the COVID-19 pandemic causing a devastating loss of life globally, dramatic changes to everyday life and significant impact on the global economy. With three novel coronaviruses emerging into humans in the last 20 years, these viruses remain an important threat to human health in the future. In addition, coronaviruses are important pathogens of animals and livestock. Infectious bronchitis virus (IBV) causes large economic losses to the UK poultry industry and there is the threat of invasion by pathogenic porcine coronaviruses into the UK.
We are studying critical virus-cell interactions required by coronaviruses to facilitate replication. Understanding how viruses interact with the host cell provides the information required for design of novel vaccine development and pan-antiviral treatment approaches and to inform surveillance in wildlife. These tools would allow better viral control, limiting economic losses, and provide preparedness for novel disease outbreaks.